Professor
Professor
Associate Dean for Research Administration
Research Integrity Officer (RIO)
Director of MEdical STudent Research Opportunities (METRO, 9/2020-6/2023)
Director of the Research Experience Program (REP, 4/2023 - ) for the 1st-year Medical Master Students in the 2-Year Program.
In 2020, Dr. Mu conceived the idea to create a students' research club and recruited 3 medical students (Jason Bard, Hemu Yeluru, and Ryan Harris) to found the EVMS Research Society (ERS).
EVMS Microbiology and Molecular Cell Biology
Dr. Mu's profile on Google Scholar, ORCiD, LinkedIn, My Bibliography (NCBI), & ResearchGate. His H index per Google Scholar: 43.
The Mu lab-created research tools deposited at Addgene have been requested 569 times by scientists worldwide.
Dr. Mu is the founding organizer of the SHARP seminars of the Leroy T. Canoles Jr. Cancer Research Center.
Office of Research, Suite 1112, Waitzer Hall, 735 Fairfax Ave, Norfolk VA 23507
Office: (757) 446-8480
Email: MuD@evms.edu
Teaching
- Biomedical Sciences Program
Education and Training
- Ph.D. – University of California at Berkeley
Adviser: Prof. Judith P. Klinman (NAS member, National Medal of Science Honoree) - Damon Runyon Postdoctoral Fellow – University of North Carolina at Chapel Hill
Adviser: Prof. Aziz Sancar (NAS, NAM member, 2015 Nobel Laureate in Chemistry)
Professional Positions
2012-Present (Tenured, 2018)
- Eastern Virginia Medical School, Norfolk, VA.
Department of Microbiology & Molecular Cell Biology
2008 - 2012
- Associate Professor
- Penn State University College of Medicine, Hershey, PA.
Departments of Pathology and Biochemistry and Molecular Biology
2004 - 2008
- Research Investigator
Cold Spring Harbor Laboratory, Cold Spring Harbor, NY.
1998 - 2004
- Scientist
The Cancer Genomics Division, Tularik Inc.
Dr. Mu has reviewed cancer research grant applications for these agencies:
- Cancer Research UK
- Lung Cancer Research Program, CDMRP, Department of Defense
- EVMS Internal Funding Mechanisms (various)
- Florida Department of Health
- Italian Ministry of Health
- Memorial Sloan Kettering Cancer Center
- National Cancer Institute
- National Institutes of Health (Dr. Mu has served 50 times as a NCI/NIH study section member)
- Ohio Cancer Research
- The Commonwealth Research Commercialization Fund (CRCF), Virginia
Dr. Mu has reviewed/edited 232 manuscripts for these 24 biomedical research journals:
- American Journal of Pathology, BMC Cancer, BBA-Mol Basis of Diseases, BioMed Research International
- British Journal of Cancer, Cancer Biology and Therapy, Cancer Research
- Chemical Research in Toxicology, Current Cancer Drug Targets, Cell Cycle
- Disease Models & Mechanisms, European Thyroid Journal, International Journal of Molecular Sciences
- Journal of Molecular Diagnostics, Journal of Pathology, J. Surgical Oncology, Molecular Cancer Research
- Molecular Therapy – Nucleic Acids, Oncogene, Oncotarget, Pediatric Research, PLOS One
- Proc. Natl. Acad. Sci. USA, Scientific Reports
Former lab members Current and Past News and Activities
Research Interests
Keywords: MicroRNA biology, Cancer Research, Lung cancer, Cancer genomics, Oncogene mechanism, Oncogenic signaling, Cancer cell metabolism, Cancer drugs, CRISPR-based gene targeting, exosome, extracellular vesicles.
I. Lung Cancer Signaling, MicroRNA, and Metabolism
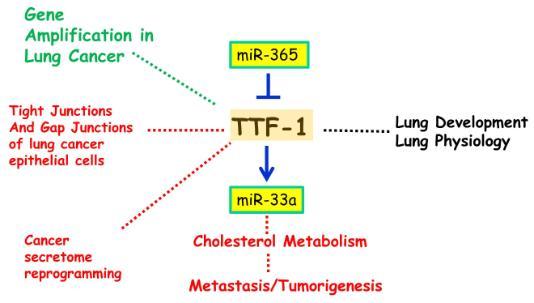
Oncogenes activated via gene amplification have a proven track record of being amenable to drug discovery. We and others discovered a recurrent amplified region in lung cancer genomes. This amplicon contains the TTF-1 gene (thyroid transcription factor 1 or known as NKX2-1) which is essential for lung development and morphogenesis. We mapped the interconnection between TTF-1 and microRNAs to uncover novel entry points to investigate TTF-1-linked lung biology and discovered the two types of microRNAs interacting with TTF-1 - miR-365 that regulates TTF-1 and miR-33a that is regulated by TTF-1. Since miR-33a is a known regulator of cholesterol metabolism (CM), our finding suggests that TTF-1 may modulate CM in the lung. This represents a novel research direction regarding TTF-1 and we are actively pursuing it.
II. Cancer Secretome and Exosome
How a cell lineage gene such as TTF-1 may reprogram the function of secretome is poorly understood. To this end, we discovered that TTF-1 confers an antiangiogenic function to the secretome of lung cancer epithelial cells. This antiangiogenic activity is derived from TTF-1-promoted secretion of GM-CSF from the lung cancer epithelial cells which in turn induces endothelial cells to release the antiangiogenic soluble VEGF receptor 1. In view of this finding, we surmise that the exosomal cargo content in the secretome may also be subject to TTF-1 regulation. Indeed, we have shown that VEGF is a direct transcriptional target of TTF-1 and the exosomal VEGF cargo level is increased in the TTF-1+ lung cancer cells, suggesting that TTF-1 may reprogram the exosomal cargo content. We are interested in quantifying the extent of the TTF-1-dependent exosomal cargo reprogramming and the associated biological consequences.
III. Junction Biology of Lung Cancer Epithelial Cells
In view of our finding that TTF-1 directly targets the expression of lung epithelial tight junction factors and the literature that TTF-1 positively regulates E-Cadherin in the adherens junction, we posit that TTF-1 may be a master regulator of the lung epithelial junction biology. To test this hypothesis, we are characterizing whether the gap junction of lung epithelial cells is also subject to TTF-1 regulation and the associated consequences of such a regulatory relationship.
Publications (My bibliography at the NCBI)
| 61. |
Bard, JT; Yeluru, H; Karpov, MV; Mu, D. Evaluating the Impact of a Medical School Student-Run Research Organization on Scholarly Activity. Cureus. 2023 Aug 7;15(8):e43067. doi: 10.7759/cureus.43067. eCollection 2023 Aug. PMID: 37680401 PMCID: PMC10481763 |
| 60. |
Martin, S; Mu, D (2022). CRISPR-Induced Loss of Connexin 43 Expression Sensitizes KRAS Mutant Cells to |
| 59. |
Fleck, AP; Flotte, AB; Si, EP; Mu, D (2022). Loss of Gap Junction Factor Connexin 43 Results in an Increase of |
| 58. |
Maria Maslyanko, Ryan D. Harris, and David Mu. Connecting Cholesterol Efflux Factors to Lung Cancer Biology and Therapeutics, International Journal of Molecular Sciences, 2021, 22: 7209. (This article belongs to the Special Issue Tumor Metabolism and Signaling) PMID: 34281263 |
| 57. |
Phelps CA, Lindsey-Boltz L, Sancar A, and Mu D. Mechanistic Study of TTF-1 Modulation of Cellular Sensitivity to Cisplatin. Scientific Reports. 2019, 9:7990. PMID: 31142791 |
| 56. |
Lai S, Phelps CA, Short AM, Sucharita DM, and Mu D. Thyroid transcription factor 1 enhances cellular statin sensitivity via perturbing cholesterol metabolism. Oncogene. 2018, 37:3290-3300. PMID: 29551766 |
| 55. |
Phelps CA, Lai S, Mu D. Roles of Thyroid Transcription Factor 1 in Lung Cancer Biology, Vitamins and Hormones (Vol. 106). Litwack G, Editor. United Kingdom, Publisher: Elsevier, 2018, pp. 517-544. (Published online in 2017). PMID: 29407447. |
| 54. |
Wood LW, Cox NI, Phelps CA, Lai SC, Poddar A, Talbot C Jr, Mu D. |
| 53. |
Maimon A, Mogilevsky M, Shilo A, Golan-Gerstl R, Obiedat A, Ben-Hur V, Lebenthal-Loinger I, Stein I, Reich R, Beenstock J, Zehorai E, Andersen CL, Thorsen K, Orntoft TF, Davis RJ, Davidson B, Mu D, Karni R. |
| 52. | Mu D. The Complexity of Thyroid Transcription Factor 1 with Both Pro- and Anti-oncogenic Activities. J Biol Chem. 2013 Aug 30;288(35):24992-5000. doi: 10.1074/jbc.R113.491647. Epub 2013 Jul 1. PubMed PMID: 23818522; PubMed Central PMCID: PMC3757165 |
| 51. | Runkle EA, Mu D. Tight junction proteins: from barrier to tumorigenesis. Cancer Lett. 2013 Aug 28;337(1):41-8. doi: 10.1016/j.canlet.2013.05.038. Epub 2013 Jun 3. PubMed PMID: 23743355; PubMed Central PMCID: PMC3752309 |
| 50. | Rice SJ, Lai SC, Wood LW, Helsley KR, Runkle EA, Winslow MM, Mu D. MicroRNA-33a mediates the regulation of high mobility group AT-hook 2 gene (HMGA2) by thyroid transcription factor 1 (TTF-1/NKX2-1). J Biol Chem. 2013 Jun 7;288(23):16348-60. doi: 10.1074/jbc.M113.474643. Epub 2013 Apr 26. PubMed PMID: 23625920; PubMed Central PMCID: PMC3675572 |
| 49. | Cohen-Eliav M, Golan-Gerstl R, Siegfried Z, Andersen CL, Thorsen K, Ørntoft TF, Mu D, Karni R. The splicing factor SRSF6 is amplified and is an oncoprotein in lung and colon cancers. J Pathol. 2013 Mar;229(4):630-9. doi: 10.1002/path.4129. PubMed PMID: 23132731 |
| 48. | Rajaram M, Zhang J, Wang T, Li J, Kuscu C, Qi H, Kato M, Grubor V, Weil RJ, Helland A, Borrenson-Dale AL, Cho KR, Levine DA, Houghton AN, Wolchok JD, Myeroff L, Markowitz SD, Lowe SW, Zhang M, Krasnitz A, Lucito R, Mu D, Powers RS. Two Distinct Categories of Focal Deletions in Cancer Genomes. PLoS One. 2013 Jun 21;8(6):e66264. Print 2013. PubMed PMID: 23805207; PubMed Central PMCID: PMC3689739 |
| 47. | Runkle EA, Rice SJ, Qi J, Masser D, Antonetti DA, Winslow MM, Mu D. Occludin is a direct target of thyroid transcription factor-1 (TTF-1/NKX2-1). J Biol Chem. 2012 Aug 17;287(34):28790-801. doi: 10.1074/jbc.M112.367987. Epub 2012 Jul 2. PubMed PMID: 22761434; PubMed Central PMCID: PMC3436544 |
| 46. | Qi J, Mu D. MicroRNAs and lung cancers: from pathogenesis to clinical implications. Front Med. 2012 Jun;6(2):134-55. doi: 10.1007/s11684-012-0188-4. Epub 2012 Apr 18. Review. PubMed PMID: 22528868; PubMed Central PMCID: PMC3725603 |
| 45. | Conkrite K, Sundby M, Mu D, Mukai S, MacPherson D. Cooperation between Rb and Arf in suppressing mouse retinoblastoma. J Clin Invest. 2012 May 1;122(5):1726-33. doi: 10.1172/JCI61403. Epub 2012 Apr 9. PubMed PMID: 22484813; PubMed Central PMCID: PMC3336990 |
| 44. | Qi J, Rice SJ, Salzberg AC, Runkle EA, Liao J, Zander DS, Mu D. MiR-365 regulates lung cancer and developmental gene thyroid transcription factor 1. Cell Cycle. 2012 Jan 1;11(1):177-86. doi: 10.4161/cc.11.1.18576. Epub 2012 Jan 1. PubMed PMID: 22185756; PubMed Central PMCID: PMC3272236 "F1000 Citation" |
| 43. | Conkrite K, Sundby M, Mukai S, Thomson JM, Mu D, Hammond SM, MacPherson D. miR-17~92 cooperates with RB pathway mutations to promote retinoblastoma. Genes Dev. 2011 Aug 15;25(16):1734-45. doi: 10.1101/gad.17027411. Epub 2011 Aug 4. PubMed PMID: 21816922; PubMed Central PMCID: PMC3165937 |
| 42. | Oliver TG, Mercer KL, Sayles LC, Burke JR, Mendus D, Lovejoy KS, Cheng MH, Subramanian A, Mu D, Powers S, Crowley D, Bronson RT, Whittaker CA, Bhutkar A, Lippard SJ, Golub T, Thomale J, Jacks T, Sweet-Cordero EA. Chronic cisplatin treatment promotes enhanced damage repair and tumor progression in a mouse model of lung cancer. Genes Dev. 2010 Apr 15;24(8):837-52. doi: 10.1101/gad.1897010. PubMed PMID: 20395368; PubMed Central PMCID: PMC2854397 |
| 41. | Hsu DS, Acharya CR, Balakumaran BS, Riedel RF, Kim MK, Stevenson M, Tuchman S, Mukherjee S, Barry W, Dressman HK, Nevins JR, Powers S, Mu D, Potti A. Characterizing the developmental pathways TTF-1, NKX2-8, and PAX9 in lung cancer. Proc Natl Acad Sci USA. 2009 Mar 31;106(13):5312-7. doi: 10.1073/pnas.0900827106. Epub 2009 Mar 11. Erratum in: Proc Natl Acad Sci U S A. 2011 Aug 30;108(35):14705. PubMed PMID: 19279207; PubMed Central PMCID: PMC2664027 |
| 40. | Sangha N, Wu R, Kuick R, Powers S, Mu D, Fiander D, Yuen K, Katabuchi H, Tashiro H, Fearon ER, Cho KR. Neurofibromin 1 (NF1) defects are common in human ovarian serous carcinomas and co-occur with TP53 mutations. Neoplasia. 2008 Dec;10(12):1362-72, following 1372. PubMed PMID: 19048115; PubMed Central PMCID: PMC2586687 |
| 39. | Miao J, Mu D, Ergel B, Singavarapu R, Duan Z, Powers S, Oliva E, Orsulic S. Hepsin colocalizes with desmosomes and induces progression of ovarian cancer in a mouse model. Int J Cancer. 2008 Nov 1;123(9):2041-7. doi: 10.1002/ijc.23726. PubMed PMID: 18726901; PubMed Central PMCID: PMC2653430 |
| 38. | Powers S, Mu D. Genetic similarities between organogenesis and tumorigenesis of the lung. Cell Cycle. 2008 Jan 15;7(2):200-4. Epub 2007 Nov 5. Review. PubMed PMID: 18256532 |
| 37. | Kendall J, Liu Q, Bakleh A, Krasnitz A, Nguyen KC, Lakshmi B, Gerald WL, Powers S, Mu D. Oncogenic cooperation and coamplification of developmental transcription factor genes in lung cancer. Proc Natl Acad Sci USA. 2007 Oct 16;104(42):16663-8. Epub 2007 Oct 9. PubMed PMID: 17925434; PubMed Central PMCID: PMC2034240 |
| 36. | Scott CL, Gil J, Hernando E, Teruya-Feldstein J, Narita M, Martínez D, Visakorpi T, Mu D, Cordon-Cardo C, Peters G, Beach D, Lowe SW. Role of the chromobox protein CBX7 in lymphomagenesis. Proc Natl Acad Sci USA. 2007 Mar 27;104(13):5389-94. Epub 2007 Mar 20. PubMed PMID: 17374722; PubMed Central PMCID: PMC1828941 |
| 35. | Karni R, de Stanchina E, Lowe SW, Sinha R, Mu D, Krainer AR. The gene encoding the splicing factor SF2/ASF is a proto-oncogene. Nat Struct Mol Biol. 2007 Mar;14(3):185-93. Epub 2007 Feb 18. PubMed PMID: 17310252 |
| 34. | MacPherson D, Conkrite K, Tam M, Mukai S, Mu D, Jacks T. Murine bilateral retinoblastoma exhibiting rapid-onset, metastatic progression and N-myc gene amplification. EMBO J. 2007 Feb 7;26(3):784-94. Epub 2007 Jan 18. PubMed PMID: 17235288; PubMed Central PMCID: PMC1794380 |
| 33. | Pelham RJ, Rodgers L, Hall I, Lucito R, Nguyen KC, Navin N, Hicks J, Mu D, Powers S, Wigler M, Botstein D. Identification of alterations in DNA copy number in host stromal cells during tumor progression. Proc Natl Acad Sci USA. 2006 Dec 26;103(52):19848-53. Epub 2006 Dec 13. PubMed PMID: 17167050; PubMed Central PMCID: PMC1698871 |
| 32. | Geurts AM, Collier LS, Geurts JL, Oseth LL, Bell ML, Mu D, Lucito R, Godbout SA, Green LE, Lowe SW, Hirsch BA, Leinwand LA, Largaespada DA. Gene mutations and genomic rearrangements in the mouse as a result of transposon mobilization from chromosomal concatemers. PLoS Genet. 2006 Sep 29;2(9):e156. Epub 2006 Aug 3. PubMed PMID: 17009875; PubMed Central PMCID: PMC1584263 |
| 31. | Zender L, Spector MS, Xue W, Flemming P, Cordon-Cardo C, Silke J, Fan ST, Luk JM, Wigler M, Hannon GJ, Mu D, Lucito R, Powers S, Lowe SW. Identification and validation of oncogenes in liver cancer using an integrative oncogenomic approach. Cell. 2006 Jun 30;125(7):1253-67. PubMed PMID: 16814713; PubMed Central PMCID: PMC3026384 |
| 30. | Sivertsen EA, Galteland E, Mu D, Holte H, Meza-Zepeda L, Myklebost O, Patzke S, Smeland EB, Stokke T. Gain of chromosome 6p is an infrequent cause of increased PIM1 expression in B-cell non-Hodgkin's lymphomas. Leukemia. 2006 Mar;20(3):539-42. PubMed PMID: 16437153 |
| 29. | Galteland E, Sivertsen EA, Svendsrud DH, Smedshammer L, Kresse SH, Meza-Zepeda LA, Myklebost O, Suo Z, Mu D, Deangelis PM, Stokke T. Translocation t(14;18) and gain of chromosome 18/BCL2: effects on BCL2 expression and apoptosis in B-cell non-Hodgkin's lymphomas. Leukemia. 2005 Dec;19(12):2313-23. PubMed PMID: 16193090 |
| 28. | He L, Thomson JM, Hemann MT, Hernando-Monge E, Mu D, Goodson S, Powers S, Cordon-Cardo C, Lowe SW, Hannon GJ, Hammond SM. A microRNA polycistron as a potential human oncogene. Nature. 2005 Jun 9;435(7043):828-33. PubMed PMID: 15944707 |
| 27. | Pei L, Wiser O, Slavin A, Mu D, Powers S, Jan LY, Hoey T. Oncogenic potential of TASK3 (Kcnk9) depends on K+ channel function. Proc Natl Acad Sci USA. 2003 Jun 24;100(13):7803-7. Epub 2003 Jun 2. PubMed PMID: 12782791; PubMed Central PMCID: PMC164668 |
| 26. | Mu D, Chen L, Zhang X, See LH, Koch CM, Yen C, Tong JJ, Spiegel L, Nguyen KC, Servoss A, Peng Y, Pei L, Marks JR, Lowe S, Hoey T, Jan LY, McCombie WR, Wigler MH, Powers S. Genomic amplification and oncogenic properties of the KCNK9 potassium channel gene. Cancer Cell. 2003 Mar;3(3):297-302. PubMed PMID: 12676587 |
| 25. | Mu D, Bessho T, Nechev LV, Chen DJ, Harris TM, Hearst JE, Sancar A. DNA interstrand cross-links induce futile repair synthesis in mammalian cell extracts. Mol Cell Biol. 2000 Apr;20(7):2446-54. PubMed PMID: 10713168; PubMed Central PMCID: PMC85433 |
| 24. | Zhao X, Mu D. (6-4) photolyase: light-dependent repair of DNA damage. Histol Histopathol. 1998 Oct;13(4):1179-82. Review. PubMed PMID: 9810509 |
| 23. | Bessho T, Mu D, Sancar A. Initiation of DNA interstrand cross-link repair in humans: the nucleotide excision repair system makes dual incisions 5' to the cross-linked base and removes a 22- to 28-nucleotide-long damage-free strand. Mol Cell Biol. 1997 Dec;17(12):6822-30. PubMed PMID: 9372913; PubMed Central PMCID: PMC232538 |
| 22. | Mu D, Wakasugi M, Hsu DS, Sancar A. Characterization of reaction intermediates of human excision repair nuclease. J Biol Chem. 1997 Nov 14;272(46):28971-9. PubMed PMID: 9360969 |
| 21. | Mu D, Sancar A. Model for XPC-independent transcription-coupled repair of pyrimidine dimers in humans. J Biol Chem. 1997 Mar 21;272(12):7570-3. PubMed PMID: 9065408 |
| 20. | Mu D, Tursun M, Duckett DR, Drummond JT, Modrich P, Sancar A. Recognition and repair of compound DNA lesions (base damage and mismatch) by human mismatch repair and excision repair systems. Mol Cell Biol. 1997 Feb;17(2):760-9. PubMed PMID: 9001230; PubMed Central PMCID: PMC231802 |
| 19. | Mu D, Sancar A. DNA excision repair assays. Prog Nucleic Acid Res Mol Biol. 1997;56:63-81. Review. PubMed PMID: 9187051 |
| 18. | Reardon JT, Mu D, Sancar A. Overproduction, purification, and characterization of the XPC subunit of the human DNA repair excision nuclease. J Biol Chem. 1996 Aug 9;271(32):19451-6. PubMed PMID: 8702634 |
| 17. | Zamble DB, Mu D, Reardon JT, Sancar A, Lippard SJ. Repair of cisplatin--DNA adducts by the mammalian excision nuclease. Biochemistry. 1996 Aug 6;35(31):10004-13. PubMed PMID: 8756462 |
| 16. | Kazantsev A, Mu D, Nichols AF, Zhao X, Linn S, Sancar A. Functional complementation of xeroderma pigmentosum complementation group E by replication protein A in an in vitro system. Proc Natl Acad Sci USA. 1996 May 14;93(10):5014-8. PubMed PMID: 8643521; PubMed Central PMCID: PMC39398 |
| 15. | Matsunaga T, Park CH, Bessho T, Mu D, Sancar A. Replication protein A confers structure-specific endonuclease activities to the XPF-ERCC1 and XPG subunits of human DNA repair excision nuclease. J Biol Chem. 1996 May 10;271(19):11047-50. PubMed PMID: 8626644 |
| 14. | Mu D, Hsu DS, Sancar A. Reaction mechanism of human DNA repair excision nuclease. J Biol Chem. 1996 Apr 5;271(14):8285-94. PubMed PMID: 8626523 |
| 13. | Matsunaga T, Mu D, Park CH, Reardon JT, Sancar A. Human DNA repair excision nuclease. Analysis of the roles of the subunits involved in dual incisions by using anti-XPG and anti-ERCC1 antibodies. J Biol Chem. 1995 Sep 1;270(35):20862-9. PubMed PMID: 7657672 |
| 12. | Park CH, Mu D, Reardon JT, Sancar A. The general transcription-repair factor TFIIH is recruited to the excision repair complex by the XPA protein independent of the TFIIE transcription factor. J Biol Chem. 1995 Mar 3;270(9):4896-902. PubMed PMID: 7876263 |
| 11. | Mu D, Park CH, Matsunaga T, Hsu DS, Reardon JT, Sancar A. Reconstitution of human DNA repair excision nuclease in a highly defined system. J Biol Chem. 1995 Feb 10;270(6):2415-8. PubMed PMID: 7852297 "Cited by 2015 Nobel Prize Award Document" |
| 10. | Mu D, Klinman JP. Cloning of mammalian topa quinone-containing enzymes. Methods Enzymol. 1995;258:114-22. PubMed PMID: 8524143 |
| 09. | Mu D, Bertrand-Burggraf E, Huang JC, Fuchs RP, Sancar A, Fuchs BP. Human and E.coli excinucleases are affected differently by the sequence context of acetylaminofluorene-guanine adduct. Nucleic Acids Res. 1994 Nov 25;22(23):4869-71. Erratum in: Nucleic Acids Res 1995 Feb 11;23(3):540. PubMed PMID: 7702657; PubMed Central PMCID: PMC523749 |
| 08. | Mu D, Medzihradszky KF, Adams GW, Mayer P, Hines WM, Burlingame AL, Smith AJ, Cai D, Klinman JP. Primary structures for a mammalian cellular and serum copper amine oxidase. J Biol Chem. 1994 Apr 1;269(13):9926-32. PubMed PMID: 8144587 |
| 07. | Klinman JP, Mu D. Quinoenzymes in biology. Annu Rev Biochem. 1994;63:299-344. Review. PubMed PMID: 7979241 |
| 06. | Mu D, Janes SM, Smith AJ, Brown DE, Dooley DM, Klinman JP. Tyrosine codon corresponds to topa quinone at the active site of copper amine oxidases. J Biol Chem. 1992 Apr 25;267(12):7979-82. PubMed PMID: 1569055 |
| 05. | Brown DE, McGuirl MA, Dooley DM, Janes SM, Mu D, Klinman JP. The organic functional group in copper-containing amine oxidases. Resonance Raman spectra are consistent with the presence of topa quinone (6-hydroxydopa quinone) in the active site. J Biol Chem. 1991 Mar 5;266(7):4049-51. PubMed PMID: 1900285 |
| 04. | Janes SM, Mu D, Wemmer D, Smith AJ, Kaur S, Maltby D, Burlingame AL, Klinman JP. A new redox cofactor in eukaryotic enzymes: 6-hydroxydopa at the active site of bovine serum amine oxidase. Science. 1990 May 25;248(4958):981-7. PubMed PMID: 2111581 |
| 03. | Tsing I, Mu D, Lee G, Peng S, Liu R. Preparation and Properties of Molybdenum-Pentadienyl Complexes: A Facile h5<=>h3 Reversible Interconversion for a Pentadienyl Ligands. Organometallics. 1989 ; 8:2248-2252 |
| 02. | Liu F, Mu D, Lee G, Peng S, Liu R. Preparation of Molybdenum-h3- Pentadienyl Complexes: Structural Characterization of a Delocalized Pentadienyl Ligand in Anti-h3 Geometry. Organometallics. 1989 ; 8:402-407 |
| 01. | Lee G, Peng S, Lush S, Mu D, Liu R. Reaction of iron-.eta.1-dienyl complexes with dienophiles. X-ray structures of the [4 + 2] cycloaddition adducts. Organometallics. 1988 ; 7:1155–1161 |
Patent Applications
| 1. | Amplified cancer gene hepsin. US Patent application number: 20030049645. Filed 2/12/2002. |
| 2. | AMPLIFIED ONCOGENES AND THEIR INVOLVEMENT IN CANCER. US Patent application number: 20030092042. Filed 8/27/2002. |
| 3. | AMPLIFICATION AND OVEREXPRESSION OF ONCOGENES. US Patent application number: 20040005615. Filed 5/22/2003. |
| 4. | GENE AMPLIFICATION AND OVEREXPRESSION IN CANCER. US Patent application number: 20050026194. Filed 6/15/2004. |
| 5. | KCNB: A NOVEL POTASSIUM CHANNEL PROTEIN. US Patent application number: 20080234470. Filed 1/3/2008. |
Patents Granted
| 1. | Diagnosis and treatment of cancer using mammalian pellino polypeptides and polynucleotides. United States Patent # 7,115,368, issued 10/3/2006. |
| 2. | Nucleic acid encoding KCNB potassium channel. United States Patent # 7,462,465, issued 12/9/2008. |
 Professor
Professor